Description
Session 1 — 19 June | 17:00–19:30
• Introduction, goals and evaluation
• Sequencing technologies and genomic data
• Online genomic databases
• Genomic data formats
• Quality control
• Online bioinformatics platforms
Session 2 — 26 June | 17:00–19:30
• Genome assembly: de novo and reference-based
• Genome annotation
Session 3 — 3 July | 17:00–19:30
• Quality metrics for genomics
• Students evaluation
Session 1- 11 September 17h-19h30
*Concepts of comparative genomics and pangenome analysis
*Core and accessory genomes
Session 2 – 18 September 17h-19h30
*Applications of comparative genomics in understanding microbial diversity and evolution
*Identifying orthologous genes using OrthoFinder
* Searching for orthologous genes in online databases
*Pangenome analysis
Session 3 – 25 September 17h-19h30
*Interpreting gene presence/absence matrices
*Students evaluation
Session 1 – 16 October 17h-19h30
*Concepts of molecular evolution and tree inference
*Basics of phylogenetic tree construction and interpretation
*Aligning sequences with MAFFT or MEGA
Session 2 – 23 October 17h-19h30
*Phylogenetic model selection
*Evolutionary insights from phylogenomic analyses
*Tree building
Session 3 – 30 October 17h-19h30
*Tree visualization with FigTree and iTOL
*Students evaluation
Session 1- 20 November 17h-19h30
*Principles of metagenomics
*Applications of metagenomics from environmental monitoring to Medicine
* Differences between metagenomics and metabarcoding
*Overview of microbiome analysis workflows
Session 2 – 27 November 17h-19h30
*Metagenomes assembly and annotation
*Taxonomic profiling
*Functional annotation of microbiomes and communities
Session 3 – 4 December 17h-19h30
*Community diversity analysis
*Students evaluation
Session 1 – 8 January 17h-19h30
*Understanding gene function and metabolic pathways
*Tools for functional annotation and pathway reconstruction.
*Functional annotation with eggNOG-mapper
Session 2 – 15 January 17h-19h30
*How to infer metabolic pathways
* Pathway analysis applied to individual genomes and metagenomic analysis
*Pathway visualization and mapping using KEGG Mapper
Session 3 – 22 January 17h-19h30
*Wrap-up
*Students evaluation
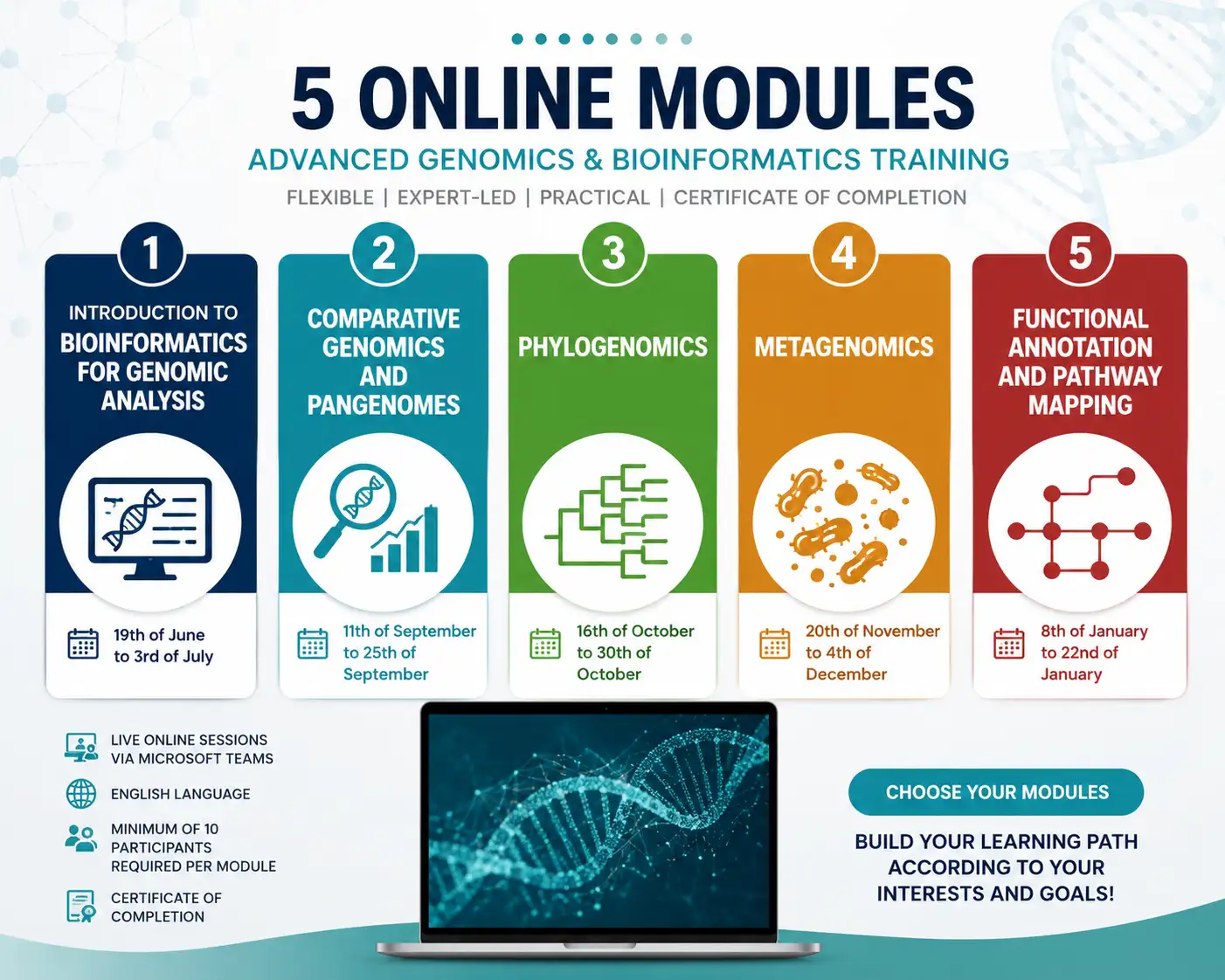
Advanced Genomics Training Series is a live online programme designed to provide participants with both foundational and advanced training in modern genomics and bioinformatics.
The rapid expansion of genomic technologies has transformed research across biology, medicine, biotechnology, agriculture, and environmental sciences. However, the ability to effectively analyse, interpret, and integrate genomic data remains a major challenge for many researchers and professionals, particularly those with limited formal training in bioinformatics or those seeking to expand their analytical capabilities into more advanced domains.
This training series provides a structured yet flexible learning pathway covering key areas of contemporary genomics, from essential bioinformatic concepts and workflows to advanced applications such as comparative genomics, pangenome analysis, phylogenomics, metagenomics, and functional annotation of genomic data.
Participants with limited prior experience will benefit from a guided introduction to sequencing technologies, genomic data formats, quality control, genome assembly, annotation, and introductory comparative analyses. The programme is designed to progressively build practical skills and conceptual understanding through real-world applications and widely used bioinformatic tools.
For participants with previous experience in genomics or bioinformatics, the course offers advanced modules focused on phylogenomic inference, comparative genomics, metagenomic workflows, and pathway reconstruction approaches. These modules combine theoretical foundations with practical applications, enabling participants to integrate modern genomic methodologies into their own research projects and analytical pipelines.
The training series is organised into independent modules, allowing participants to tailor their learning experience according to their research interests, professional background, and technical expertise. Participants may enrol in individual modules or combine multiple modules with automatic discounted pricing, including the option to complete the full training series.
All sessions are delivered live online via Microsoft Teams in English language and include opportunities for discussion, practical interpretation of genomic data, and participant evaluation.